Structural Biologist | Protein Engineer
Professor Chaithanya Madhurantakam is a structural biologist whose career is defined by a deep focus on the molecular architecture of proteins. His work spans structural immunology, protein engineering, and reverse vaccinology, utilizing X-ray crystallography, biochemical and biophysical techniques to solve complex biological puzzles.
Research Group: Structural and Molecular Biology Laboratory (SMBL), TERI SAS
At SMBL, DBT- TERI SAS, Prof. Madhurantakam’s research marks a major lead in combating pathogens through molecular characterization studies since 2014. SMBL researchers are involved in atomic and molecular level characterization studies of macromolecules in Mycobacterium tuberculosis, Helicobacter pylori and functionalization of nano-molecules. The core research capabilities of SMBL group are broadly within: (i) Infectious Diseases and Antimicrobial Discovery (ii) Structural Immunology and Reverse Vaccinology, (iii) Advanced Protein Engineering.
Current R&D activities at SMBL:
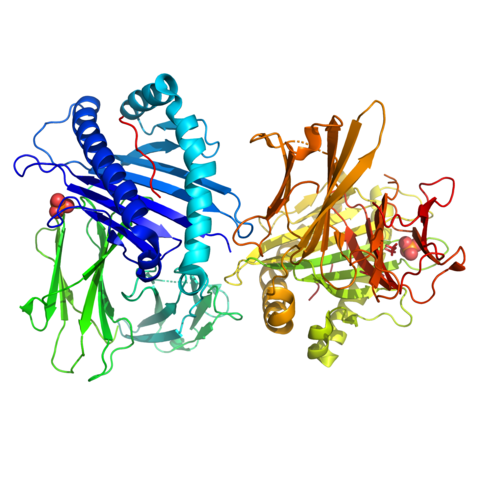
(a) The "Waxy Wall" & MmaA1: His research team provided the first atomic-level characterization of the MmaA1 enzyme of Mycobacterium tuberculosis in three states through X-ray crystallographic analysis: apo, SAM-bound, and SAH-bound.
(b) BVMOs (EthA & MymA): Using an integrated Reverse Vaccinology and CADD approach, the research team is involved in the generation of next-generation multi-epitope vaccine candidates to trigger robust immune responses against Tuberculosis.
(c) MmaA3 & Drug Screening: Utilizing the MM-GBSA method, his group screened thousands of compounds to identify inhibitors that block methoxy-mycolic acid synthesis, that may effectively weaken the bacterial defence mechanism.
(d) H. pylori Survival: The group also focuses on dUTPase and HppA in H. pylori, identifying small molecules that force the bacteria into "thymine-less death" while disrupting its energy management in acidic stomach environments.
R&D Experience:
- Structural Immunology at the Karolinska Institute
In close collaboration with Prof. Adnane Achour at the Karolinska Institute (2008-2010), Prof. Madhurantakam’s research has been instrumental in understanding immune recognition and viral subversion: (a) Altered Peptide Ligands (APLs): His work investigated how subtle changes in peptide sequences can significantly alter T-cell responses. (b) Immune Evasion: This study detailed how viruses evolve to "hide" from the host immune system by modifying their structural presentation within MHC class I complexes. (c) Allergen Research: He contributed to the structural basis of mammalian allergens working with Prof. Adnane Achour and Prof. Hans Grönlund, providing a blueprint for how the body identifies specific environmental proteins as threats.
- Protein Engineering at the University of Zurich
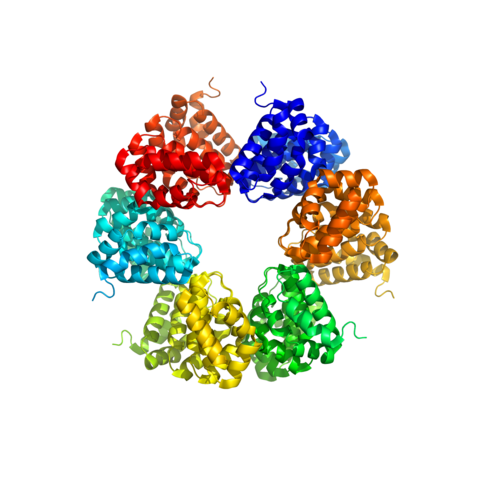
During his post-doctoral tenure at the University of Zurich (2010-2013), Prof. Madhurantakam engaged in wide-ranging collaborations involving Prof. Andreas Plückthun (Principal Investigator), Dr. Peer Mittl, Prof. Markus Grutter, and other research groups of Prof. David Baker, Prof. Zerbe, and Prof. Caflisch for the Armadillo Repeat Project: (a) dArm-Repeat Proteins (dArmRPs): This study was focused on designed Armadillo repeat proteins—modular, synthetic scaffolds engineered to bind specific peptide targets with high affinity, moving beyond traditional antibody-based approaches.
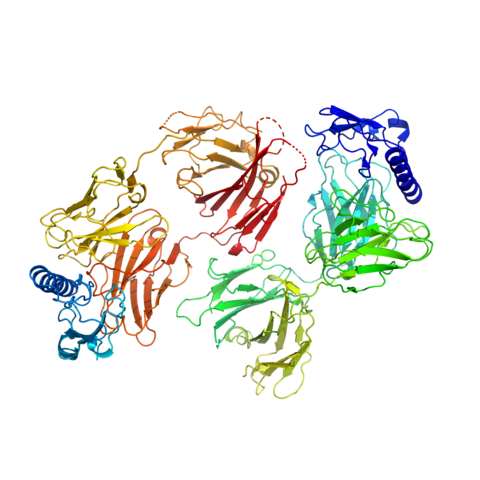
Additionally, working with Prof. Markus Grutter, MedImmune (Cambridge, UK) and AstraZeneca (Mölndal, Sweden), he was instrumental in characterizing the Gipg013 monoclonal antibody: (a) Metabolic Signalling: He helped characterize Gipg013, a highly potent monoclonal antibody. By solving the crystal structure of the Gipg013 Fab fragment in complex with the human GIPr Extracellular Domain (ECD), the study revealed how the antibody acts as a competitive antagonist by physically blocking the natural hormone binding site.
- Macromolecular Studies at IIT Kharagpur, CCMB, and Vrije Universiteit
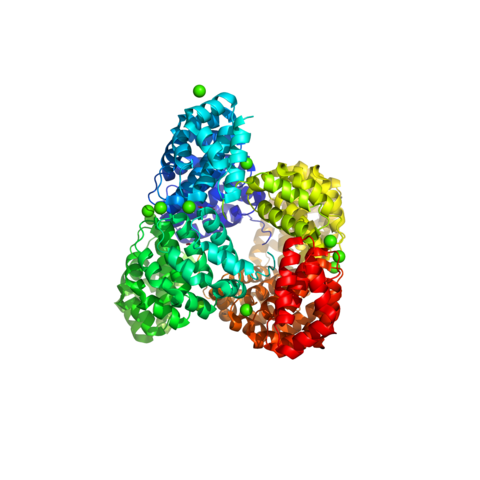
Professor Madhurantakam’s formative research involved working on the "structure-function" relationship of target molecules that are key for drug discovery strategies on following projects: (a) M. tuberculosis Pathogenesis (IIT KGP & CCMB, 2003-2008): Working with Prof. Amit Kumar Das and Dr. Rajan Sankaranarayanan, he solved the structures of essential proteins such as LMWPTPase- MPtpA, identifying critical "weak spots" in Mtb metabolic pathways. (b) Surface Layer Proteins (SLPs) (Vrije Universiteit Brussel, 2013-2014): Collaborating with Prof. Han Remaut, he explored the self-assembling world of S-layers—the proteinaceous coats that cover the cell surfaces of bacteria like Bacillus.
Research Publications -Pubmed Indexed (with 2-year IF):
27. Kaur S, Sisodia R, Pareek M, Sri T, Chattopadhyay S, Yaduvanshi P, Asthana S, Madhurantakam C, Singh A. Molecular dynamics and evolutionary conservation of FUL-SOC1 regulatory interactions in polyploid Brassica juncea. Funct Integr Genomics. 2026;26(1):67. doi:10.1007/s10142-026-01831-1 (IF: 3.0)
26. Chaudhary B, Kobakhidze G, Wachelder L, Mazumdar PA, Dong G, Madhurantakam C. Crystal structures of the mycolic acid methyl transferase 1 (MmaA1) from Mycobacterium tuberculosis in the apo-form and in complex with different cofactors reveal unique features for substrate binding. J Biomol Struct Dyn. Published online April 9, 2025:1-10. doi:10.1080/07391102.2025.2483952 (IF: 2.9)
25. Kaur S, Sisodia R, Gupta B, Gaikwad K, Madhurantakam C, Singh A. Multiple combinatorial interactions among natural structural variants of Brassica SOC1 promoters and SVP: conservation of binding affinity despite diversity in bimolecular interactions. Mol Biol Rep. 2025;52(1):187. doi:10.1007/s11033-024-10182-8 (IF: 2.7)
24. Chaudhary B, Sisodia R, Sarmadhikari D, Mazumdar PA, Asthana S, Madhurantakam C. In silico analysis and characterization of potential inhibitors of MmaA3, a methoxy mycolic acid synthase from Mycobacterium tuberculosis. J Biomol Struct Dyn. 2025;43(14):7771-7796. doi:10.1080/07391102.2024.2349545 (IF: 2.9)
23. Sisodia R, Sarmadhikari D, Mazumdar PA, Asthana S, Madhurantakam C. Molecular analysis of dUTPase of Helicobacter pylori for identification of novel inhibitors using in silico studies. J Biomol Struct Dyn. 2024;42(16):8598-8623. doi:10.1080/07391102.2023.2247080 (IF: 2.9)
22. Sisodia R, Mazumdar PA, Madhurantakam C. In silico identification and analysis of potential inhibitors for acid phosphatase, HppA from Helicobacter pylori. J Mol Recognit. 2023;36(9):e3049. doi:10.1002/jmr.3049 (IF: 3.0)
21. Suthar S, Das S, Nagpure A, Madhurantakam C, Tiwari SB, Gahlot P, Tyagi VK. Epidemiology and diagnosis, environmental resources quality and socio-economic perspectives for COVID-19 pandemic. J Environ Manage. 2021;280:111700. doi:10.1016/j.jenvman.2020.111700 (IF: 8.4)
20. Duru AD, Sun R, Allerbring EB, et al. Tuning antiviral CD8 T-cell response via proline-altered peptide ligand vaccination. PLoS Pathog. 2020;16(5):e1008244. doi:10.1371/journal.ppat.1008244 (IF: 5.0)
19. Tyagi S, Mazumdar PA, Mayee P, et al. Natural variation in Brassica FT homeologs influences multiple agronomic traits including flowering time, silique shape, oil profile, stomatal morphology and plant height in B. juncea. Plant Sci. 2018;277:251-266. doi:10.1016/j.plantsci.2018.09.018 (IF: 4.1)
18. Hansen S, Kiefer JD, Madhurantakam C, Mittl PRE, Plückthun A. Structures of designed armadillo repeat proteins binding to peptides fused to globular domains. Protein Sci. 2017;26(10):1942-1952. doi:10.1002/pro.3229 (IF: 8.0)
17. Reichen C, Hansen S, Forzani C, et al. Computationally Designed Armadillo Repeat Proteins for Modular Peptide Recognition. J Mol Biol. 2016;428(22):4467-4489. doi:10.1016/j.jmb.2016.09.012 (IF: 4.5)
16. Reichen C, Madhurantakam C, Hansen S, Grütter MG, Plückthun A, Mittl PR. Structures of designed armadillo-repeat proteins show propagation of inter-repeat interface effects. Acta Crystallogr D Struct Biol. 2016;72(Pt 1):168-175. doi:10.1107/S2059798315023116 (IF: 3.8)
15. Hansen S, Tremmel D, Madhurantakam C, Reichen C, Mittl PR, Plückthun A. Structure and Energetic Contributions of a Designed Modular Peptide-Binding Protein with Picomolar Affinity. J Am Chem Soc. 2016;138(10):3526-3532. doi:10.1021/jacs.6b00099 (IF: 15.7)
14. Reichen C, Madhurantakam C, Plückthun A, Mittl PR. Crystal structures of designed armadillo repeat proteins: implications of construct design and crystallization conditions on overall structure. Protein Sci. 2014;23(11):1572-1583. doi:10.1002/pro.2535 (IF: 8.0)
13. Ravn P, Madhurantakam C, Kunze S, et al. Structural and pharmacological characterization of novel potent and selective monoclonal antibody antagonists of glucose-dependent insulinotropic polypeptide receptor. J Biol Chem. 2013;288(27):19760-19772. doi:10.1074/jbc.M112.426288 (IF: 4.0)
12. Allerbring EB, Duru AD, Uchtenhagen H, Madhurantakam C, et al. Unexpected T-cell recognition of an altered peptide ligand is driven by reversed thermodynamics. Eur J Immunol. 2012;42(11):2990-3000. doi:10.1002/eji.201242588 (IF: 5.5)
11. Madhurantakam C, Varadamsetty G, Grütter MG, Plückthun A, Mittl PR. Structure-based optimization of designed Armadillo-repeat proteins. Protein Sci. 2012;21(7):1015-1028. doi:10.1002/pro.2085 (IF: 8.0)
10. Madhurantakam C, Duru AD, Sandalova T, Webb JR, Achour A. Inflammation-associated nitrotyrosination affects TCR recognition through reduced stability and alteration of the molecular surface of the MHC complex. PLoS One. 2012;7(3):e32805. doi:10.1371/journal.pone.0032805 (IF: 2.9)
9. Madhurantakam C, Nilsson OB, Uchtenhagen H, et al. Crystal structure of the dog lipocalin allergen Can f 2: implications for cross-reactivity to the cat allergen Fel d 4. J Mol Biol. 2010;401(1):68-83. doi:10.1016/j.jmb.2010.05.043 (IF: 4.5)
8. Neiers F, Madhurantakam C, Fälker S, et al. Two crystal structures of pneumococcal pilus sortase C provide novel insights into catalysis and substrate specificity. J Mol Biol. 2009;393(3):704-716. doi:10.1016/j.jmb.2009.08.058 (IF: 4.5)
7. Uchtenhagen H, Sourial S, Friemann R, et al. Production, purification, crystallization and preliminary X-ray diffraction analysis of the HIV-2-neutralizing V3 loop-specific Fab fragment 7C8. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009;65(Pt 7):705-708. doi:10.1107/S1744309109020685 (IF: 0.7)
6. Madhurantakam C, Nilsson OB, Jönsson K, Grönlund H, Achour A. Production, crystallization and preliminary X-ray diffraction analysis of the allergen Can f2 from Canis familiaris. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009;65(Pt 5):467-471. doi:10.1107/S1744309109010884 (IF: 0.7)
5. Roy S, Aravind P, Madhurantakam C, Ghosh AK, Sankaranarayanan R, Das AK. Crystal structure of a fungal protease inhibitor from Antheraea mylitta. J Struct Biol. 2009;166(1):79-87. doi:10.1016/j.jsb.2008.12.010 (IF: 2.7)
4. Chavali VR, Madhurantakam C, Ghorai S, Roy S, Das AK, Ghosh AK. Genome segment 6 of Antheraea mylitta cypovirus encodes a structural protein with ATPase activity. Virology. 2008;377(1):7-18. doi:10.1016/j.virol.2008.03.038 (IF: 3.8)
3. Madhurantakam C, Chavali VR, Das AK. Analyzing the catalytic mechanism of MPtpA: a low molecular weight protein tyrosine phosphatase from Mycobacterium tuberculosis through site-directed mutagenesis. Proteins. 2008;71(2):706-714. doi:10.1002/prot.21816 (IF: 2.8)
2. Roy S, Aravind P, Madhurantakam C, et al. Crystallization and preliminary X-ray diffraction analysis of a protease inhibitor from the haemolymph of the Indian tasar silkworm Antheraea mylitta. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2006;62(Pt 7):669-671. doi:10.1107/S1744309106020719 (IF: 0.7)
1. Madhurantakam C, Rajakumara E, Mazumdar PA, et al. Crystal structure of low-molecular-weight protein tyrosine phosphatase from Mycobacterium tuberculosis at 1.9-A resolution. J Bacteriol. 2005;187(6):2175-2181. doi:10.1128/JB.187.6.2175-2181.2005 (IF: 2.8)
Invited Chapters/ Reviews (2)
1: Madhurantakam, C; Howorka, S; Remaut, H. Involvement of SbsB in Ca2+ triggered S-layer assembly. Encyclopedia of Inorganic and Bioinorganic Chemistry (EIBC), John Wiley & Sons. 2014
2: Madhurantakam C; Howorka, S; Remaut H. Structure of S-layer in bacteria and archaea. Nanomicrobiology: Physiological and Environmental Charactersitics (Springer). 2014
Proceedings published in conferences and seminars (4)
1: Allerbring EB, Duru AD, Chadderton J, Markov N, Uchtenhagen H, Popov A, Madhurantakam C, Sandalova T, Turner SJ, Nygren PÅ, Achour A. Structural and thermodynamic basis underlying in vivo reestablishment of T-cell recognition of a viral escape mutant. Molecular Immunology. 68(2) 151-151, 2015. (IF: 3.4)
2: Hansen S, Reichen C, Madhurantakam C, Grutter M, Pluckthun A, Mittl PRE. Structure based engineering of designed armadillo repeat proteins. Acta Cryst A70, A70, C1153, 2014, (IF: 2.33)
3: Reichen, C; Varadamsetty, G; Tremmel, D; Parmeggiani, F; Honegger, A; Fleishmann, S; Baker, D; Madhurantakam, C; Mittl, P; Grutter, M; Pluckthun, A. Design and Selection of Armadillo repeat proteins: A novel technology for modular peptide recognition. Protein Science, 21, 82, 2012. Conference: 26th Annual Symposium of the Protein-Society, San Diego, CA, AUG-05-08, 2012. (IF: 8.0)
4: Allerbring, EB; Duru, AD; Uchtenhagen, H; Madhurantakam, C; Tomek, MB; Grimm, S; Mazumdar, PA; Friemann, R; Sandalova, T; Uhlin, M; Nygren, PA; Achour, A. The unexpected T-cell recognition of an altered peptide ligand is driven by reversed thermodynamics and an alternative structural hotspot. Immunology, 70, 2011. (IF: 5.0)